A sigma factor (σ factor) (specificity factor) is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. It is homologous to archaeal transcription factor B and to eukaryotic factor TFIIB..
Also asked, is sigma factor only in prokaryotes?
Sigma confers upon the RNA polymerase holoenzyme the ability to initiate RNA synthesis at specific locations in DNA called promoters. To bind promoter-specific regions, the core enzyme requires another subunit, sigma (σ). All prokaryotes contain multiple sigma factors.
One may also ask, how does sigma factors recognize promoter? Alternative Sigma Factors. Sigma factors act by binding to and influencing the promoter specificity of the RNA polymerase core enzyme, thereby directing selective transcription of different gene sets, coordinating gene expression in response to various environmental and endogenous cues.
One may also ask, what are alternative sigma factors?
Abstract. Alternative sigma factors enable bacteria to change the promoter specificity of the core RNA polymerase to enable the expression of genes that give them advantages in particular situations. The number of alternative sigma factors that bacteria produce varies greatly.
Why does E coli have several different sigma factors?
A) They Allow Different RNA Polymerases To Bind To The Promoters B) They Allow The Different Subunits Of The RNA Polymerase Holoenzyme To Bind To Each Other C) There Is No Good Reason, They All Perform The Same Function D) One Is Needed To Transcribe MRNA.
Related Question Answers
How does sigma factor work?
A sigma factor (σ factor) (specificity factor) is a protein needed for initiation of transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to gene promoters. The number of sigma factors varies between bacterial species.Are sigma factors transcription factors?
Sigma factors (sigmas) are bacterial transcription factors that bind core RNA polymerase (RNAP) and direct transcription initiation at cognate promoter sites. require the assistance of protein transcription factors to start synthesis.What RNA polymerase is used in prokaryotic transcription?
Prokaryotes utilize one RNA polymerase for all transcription of types of RNA. In contrast, eukaryotes utilize three slightly different RNA polymerases: RNA polymerase I, RNA polymerase II, and RNA polymerase III (8). Each of the three RNA polymerases in eukaryotes is responsible for transcribing a unique type of RNA.Which region contains the promoter?
A promoter is a regulatory region of DNA located upstream (towards the 5' region) of of a gene, providing a control point for regulated gene transcription. The promoter contains specific DNA sequences that are recognized by proteins known as transcription factors.Is the TATA box transcribed?
Transcription is a process that produces an RNA molecule from a DNA sequence. The TATA box is named for its conserved DNA sequence, which is most commonly TATAAA. Many eukaryotic genes have a conserved TATA box located 25-35 base pairs before the transcription start site of a gene.Where does transcription occur in the cell?
In a prokaryotic cell, transcription and translation are coupled; that is, translation begins while the mRNA is still being synthesized. In a eukaryotic cell, transcription occurs in the nucleus, and translation occurs in the cytoplasm.What does Rho protein do?
Rho protein functions as a hexamer of a single polypeptide chain with 419 residues, which is the product of the rho gene. It is an RNA-binding protein with the capacity to hydrolyze ATP and other nucleoside triphosphates.Is RNA polymerase a Holoenzyme?
RNA polymerase II holoenzyme is a form of eukaryotic RNA polymerase II that is recruited to the promoters of protein-coding genes in living cells. It consists of RNA polymerase II, a subset of general transcription factors, and regulatory proteins known as SRB proteins.Do eukaryotes have operons?
Operons occur in prokaryotes, but not eukaryotes. In eukaryotes, each gene is made on individual mRNAs and each gene has its own promoter. Operons are prokaryotic arrangements of multiple genes (with common functions) under the control of a single promoter.What is core polymerase?
From Wikipedia, the free encyclopedia. A core enzyme consists of the subunits of an enzyme that are needed for catalytic activity, as in the core enzyme RNA polymerase. An example of a core enzyme is a RNA polymerase enzyme without the sigma factor (σ).Is RNA polymerase a transcription factor?
RNA polymerase can attach to the promoter only with the help of proteins called basal (general) transcription factors. They are part of the cell's core transcription toolkit, needed for the transcription of any gene.What is the direction of synthesis of RNA?
RNA polymerase synthesizes an RNA strand complementary to a template DNA strand. It synthesizes the RNA strand in the 5' to 3' direction, while reading the template DNA strand in the 3' to 5' direction. The template DNA strand and RNA strand are antiparallel.Is the terminator sequence transcribed?
The terminator is a region of DNA that includes the sequence that codes for the Rho binding site in the mRNA, as well as the actual transcription stop point (which is a sequence that causes the RNA polymerase to pause so that Rho can catch up to it).What is DNA dependent RNA polymerase?
showAvailable protein structures: RNA-dependent RNA polymerase (RdRP), (RDR), or RNA replicase, is an enzyme that catalyzes the replication of RNA from an RNA template. This is in contrast to a typical DNA-dependent RNA polymerase, which catalyzes the transcription of RNA from a DNA template.Do prokaryotes have transcription factors?
In prokaryotes, transcription and translation can occur simultaneously in the cytoplasm of the cell, whereas in eukaryotes transcription occurs in the nucleus and translation occurs in the cytoplasm. Prokaryotes have a σ-factor that detects and binds to promoter sites but eukaryotes do not need a σ-factor.What is a promoter site?
promoter. Promoter sequences are DNA sequences that define where transcription of a gene by RNA polymerase begins. Promoter sequences are typically located directly upstream or at the 5' end of the transcription initiation site.What is the role of the sigma subunit of RNA polymerase?
The sigma (sigma) subunit of eubacterial RNA polymerase enables the holoenzyme to recognize and bind to specific sites on DNA known as promoters. Bacteria from several general possess multiple sigma factors that enable RNA polymerase to utilize difference classes of promoters.What is the synthesis of mRNA called?
mRNA is synthesized in the nucleus using the nucelotide sequence of DNA as a template. This process requires nucleotide triphosphates as substrates and is catalyzed by the enzyme RNA polymerase II. The process of making mRNA from DNA is called transcription, and it occurs in the nucleus.What protein recognizes promoters in bacteria?
The most abundant σ factor in the bacterial cell, referred to as σ70 inE. coli(σA in most other species), is responsible for recognizing most of the cell's promoters. σ70 contains four regions of conserved amino acid sequences, σ1–σ4, that are further divided into subregions1 (see the figure). 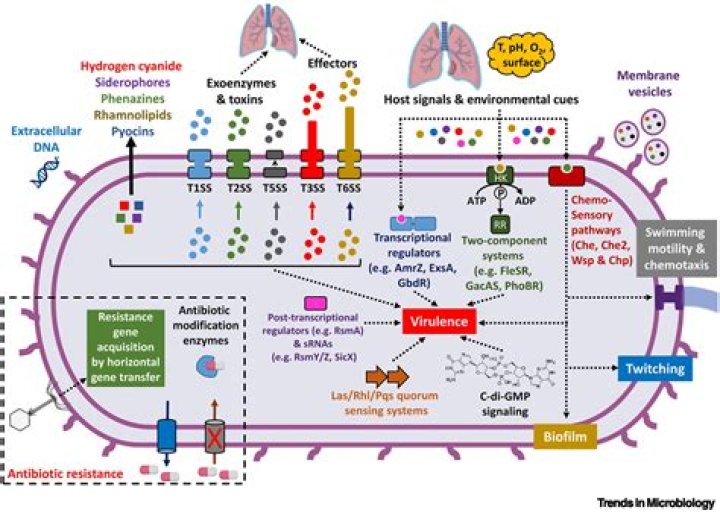
 Isabella Bartlett
Isabella Bartlett